As the food industry is moving toward a more preventive food safety strategy, environmental monitoring is playing an increasingly critical role in testing. Hazard analysis is shifting the focus from finished product testing to proactively testing the environment and the processing as critical control points to continuously monitor and reduce risk. Today many facilities are adding or strengthening their environmental monitoring programs to enhance their food safety risk reduction efforts.
In a recent webinar, Ann Draughon, Emeritus Professor of Food Microbiology and Toxicology, University of Tennessee spoke about Developing an Effective Environmental Monitoring, Sampling and Testing (EMS) Program. We present some excerpts from her presentation.
What do you need to get started with an EMS program?
“You need to first identify the right team; think about what kind of food you are processing (raw products or ready-to-eat products) and if it has had any food safety outbreak associated with it; determine critical or hygiene zones in your facility; determine sample locations; finalize which indicator tests will be done, and in which zones; determine which pathogens you will test for; choose the right test methods; set a baseline, and link that with your sampling plan, and establish testing frequency once you have finalized the number of samples and zones,” explains Draughon.
To establish critical hygiene zones, she advises to:
- Survey entire facility and have a map of that facility;
- Study that map and identify traffic patterns to divide the facility into critical hygiene zones, GMP zones, and non-processing zones;
- Put in place barriers between these zones and dedicate equipment to the critical hygiene zone, and restrict access between zones; and
- Establish strict cleaning, sanitation and monitoring plans for these diff zones.
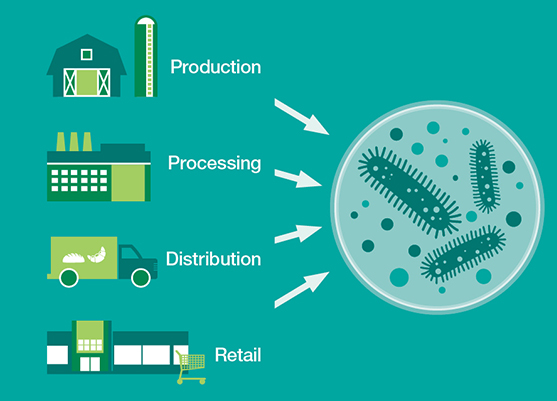
Sampling of zones should be based on risk of contamination and/ or transmission of pathogens to food from environment, says Draughon. The sampling should also take into account potential sources of product contamination by whatever means during food processing (see image 1 for examples of 4 zone and 3-zone hygiene systems).
Selecting the right assays for your EMS program
There are many options, and it can be confusing to select the right assay for your needs. Draughon advises that companies need to look their monitoring needs and consider both indicator bacteria and pathogenic bacteria to select the right assay.
For monitoring with indicator bacteria, most companies look at ATP for environmental sanitation, often before start-up to make sure facility is clean before processing begins. Protein assays are also used to pick up any allergen on equipment.
APC or total viable count is a simple assay offering many choices, which tests for the number of live bacteria on your equipment or in your environment that can grow under air or oxygen at room temperature.
Yeast/ mold count assays are good for two purposes: 1. Mold frequently is the cause of spoilage in food, so it’s useful to understand if there are any present to determine shelf life, and 2. It also helps us understand the number of particulates in the air.
We can also select specific microbial groups as indicators, such as total Enterobacteriacae, fecal coliform or E.coli or Listeria species.
Sample collection and prep
When we collect a sample, we have to clearly document the sample including information such as when it was taken, from where, by whom, what happened to that sample etc. Use clean SOPs to reduce error. Use the assays previously selected and do it as quickly as feasible. If you are working with an outside company, decide how they are going to handle the sample. Finally, always keep in mind plant safety and leave nothing behind after sampling, and avoid cross-contamination.
For characterizing pathogens, you may want to genetically fingerprint any pathogenic isolates from your facilities. This will allow you to see if you have a constant harborage of a particular pathogen or if it changes. Draughon recommends using a contract lab for characterizing pathogens, as they would be better suited, and have better resources to do this. Destroy the isolates after characterization – you don’t want any chance of the pathogen spreading into the product or the environment.
Written SOPs for EMS programs
It’s critical to have clear written SOPs for EMS programs which include the following:
- Frequency of sampling;
- When, where , how and duration of sampling;
- Procedure for recording data and coding;
- Sample number, size or volume;
- Specific sampling and analysis validated protocols;
- Monitoring of incubators and use of equipment;
- Handling and shipping of samples; and
- Alert and action levels and appropriate response to deviations from alert or action levels.
It’s also important that we train and validate the personnel performing EMS. Each individual doing this needs to demonstrate proficiency of doing this. They need to understand proper recording of EMS program data, alert and action levels, and zero tolerance levels. The personnel should be comfortable and qualified for sampling protocol, and using all the equipment.
In summary, sampling plans should be adaptable, which highest risk sites being tested initially. Establish a baseline and modify sampling plan as needed. Establish your sampling and testing criteria and sample as needed with each zone to fully assess the environmental program.
 One of the goals of the consortium is to see if there are any actions that food producers can take in respect to microbiomes that can reduce risk and make production safer. And Mars, with more than 130 factories worldwide, can help map the flow of microorganisms into and through the supply chain on a global level. An informatics infrastructure developed in the IBM Accelerated Discovery Lab, a data and analytics hub for IBM researchers and their clients and partners, will help the team parse and aggregate terabytes of genomic data from Mars and apply decades of refined analytics to uncover new insights. Adding relevant weather, transport and other contextual data could help define a targeted breakout, marking on the index a warning for food producers and distributors at the outset of a processing cycle.
One of the goals of the consortium is to see if there are any actions that food producers can take in respect to microbiomes that can reduce risk and make production safer. And Mars, with more than 130 factories worldwide, can help map the flow of microorganisms into and through the supply chain on a global level. An informatics infrastructure developed in the IBM Accelerated Discovery Lab, a data and analytics hub for IBM researchers and their clients and partners, will help the team parse and aggregate terabytes of genomic data from Mars and apply decades of refined analytics to uncover new insights. Adding relevant weather, transport and other contextual data could help define a targeted breakout, marking on the index a warning for food producers and distributors at the outset of a processing cycle.